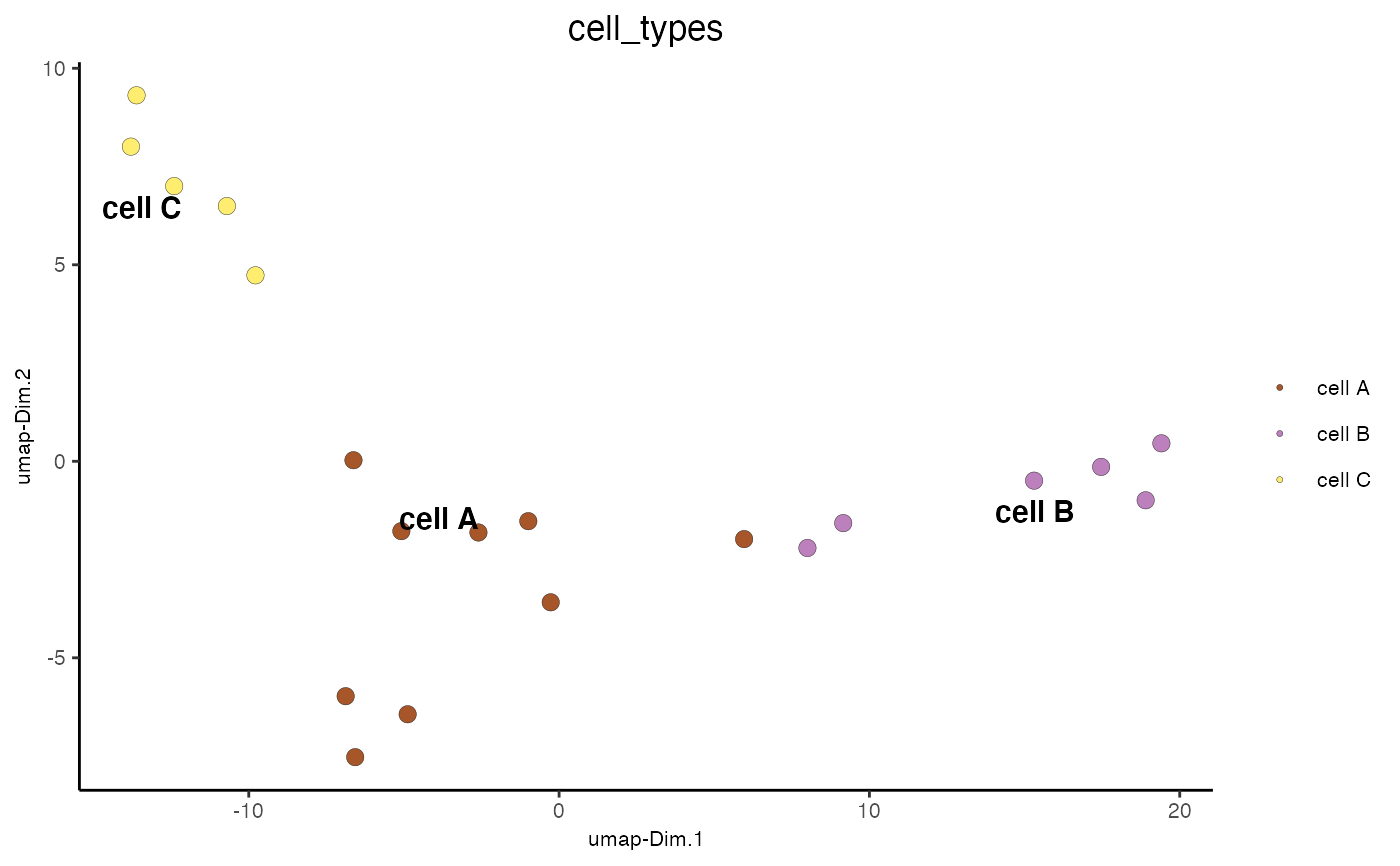
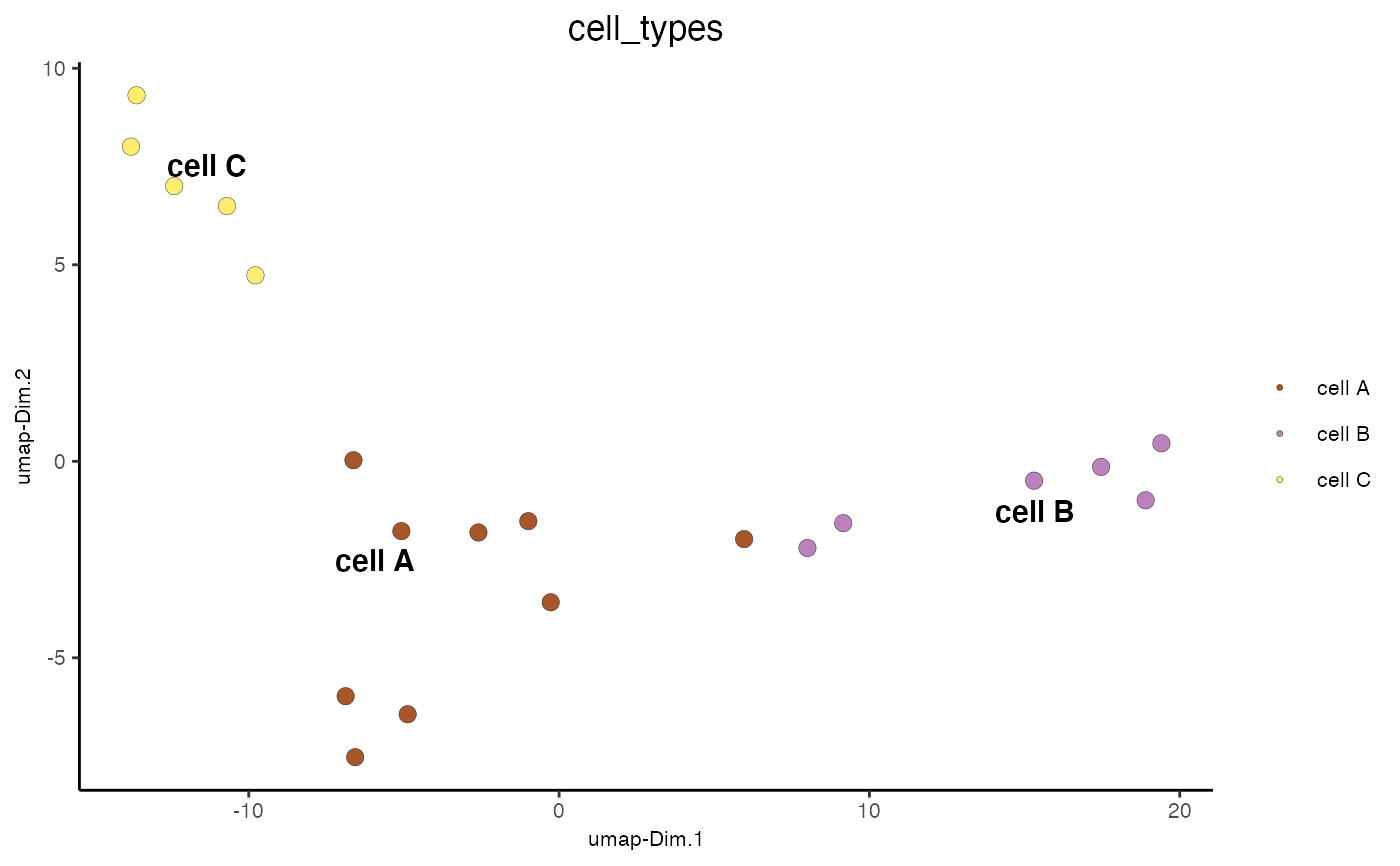
Visualize cells according to dimension reduction coordinates
dimPlot2D( gobject, group_by = NULL, group_by_subset = NULL, dim_reduction_to_use = "umap", dim_reduction_name = "umap", dim1_to_use = 1, dim2_to_use = 2, spat_enr_names = NULL, show_NN_network = F, nn_network_to_use = "sNN", network_name = "sNN.pca", cell_color = NULL, color_as_factor = T, cell_color_code = NULL, cell_color_gradient = c("blue", "white", "red"), gradient_midpoint = NULL, gradient_limits = NULL, select_cell_groups = NULL, select_cells = NULL, show_other_cells = T, other_cell_color = "lightgrey", other_point_size = 0.5, show_cluster_center = F, show_center_label = T, center_point_size = 4, center_point_border_col = "black", center_point_border_stroke = 0.1, label_size = 4, label_fontface = "bold", edge_alpha = NULL, point_shape = c("border", "no_border"), point_size = 1, point_alpha = 1, point_border_col = "black", point_border_stroke = 0.1, title = NULL, show_legend = T, legend_text = 8, legend_symbol_size = 1, background_color = "white", axis_text = 8, axis_title = 8, cow_n_col = 2, cow_rel_h = 1, cow_rel_w = 1, cow_align = "h", show_plot = NA, return_plot = NA, save_plot = NA, save_param = list(), default_save_name = "dimPlot2D" )
Arguments
| gobject | giotto object |
|---|---|
| group_by | create multiple plots based on cell annotation column |
| group_by_subset | subset the group_by factor column |
| dim_reduction_to_use | dimension reduction to use |
| dim_reduction_name | dimension reduction name |
| dim1_to_use | dimension to use on x-axis |
| dim2_to_use | dimension to use on y-axis |
| spat_enr_names | names of spatial enrichment results to include |
| show_NN_network | show underlying NN network |
| nn_network_to_use | type of NN network to use (kNN vs sNN) |
| network_name | name of NN network to use, if show_NN_network = TRUE |
| cell_color | color for cells (see details) |
| color_as_factor | convert color column to factor |
| cell_color_code | named vector with colors |
| cell_color_gradient | vector with 3 colors for numeric data |
| gradient_midpoint | midpoint for color gradient |
| gradient_limits | vector with lower and upper limits |
| select_cell_groups | select subset of cells/clusters based on cell_color parameter |
| select_cells | select subset of cells based on cell IDs |
| show_other_cells | display not selected cells |
| other_cell_color | color of not selected cells |
| other_point_size | size of not selected cells |
| show_cluster_center | plot center of selected clusters |
| show_center_label | plot label of selected clusters |
| center_point_size | size of center points |
| center_point_border_col | border color of center points |
| center_point_border_stroke | border stroke size of center points |
| label_size | size of labels |
| label_fontface | font of labels |
| edge_alpha | column to use for alpha of the edges |
| point_shape | point with border or not (border or no_border) |
| point_size | size of point (cell) |
| point_alpha | transparancy of point |
| point_border_col | color of border around points |
| point_border_stroke | stroke size of border around points |
| title | title for plot, defaults to cell_color parameter |
| show_legend | show legend |
| legend_text | size of legend text |
| legend_symbol_size | size of legend symbols |
| background_color | color of plot background |
| axis_text | size of axis text |
| axis_title | size of axis title |
| cow_n_col | cowplot param: how many columns |
| cow_rel_h | cowplot param: relative height |
| cow_rel_w | cowplot param: relative width |
| cow_align | cowplot param: how to align |
| show_plot | show plot |
| return_plot | return ggplot object |
| save_plot | directly save the plot [boolean] |
| save_param | list of saving parameters, see |
| default_save_name | default save name for saving, don't change, change save_name in save_param |
Value
ggplot
Details
Description of parameters. For 3D plots see dimPlot3D
See also
Other reduced dimension visualizations:
dimPlot3D(),
dimPlot(),
plotPCA_2D(),
plotPCA_3D(),
plotPCA(),
plotTSNE_2D(),
plotTSNE_3D(),
plotTSNE(),
plotUMAP_2D(),
plotUMAP_3D(),
plotUMAP()
Examples
dimPlot2D(mini_giotto_single_cell, cell_color = 'cell_types', point_size = 3)